TIDER: Tracking of Insertion, DEletions and Recombination events
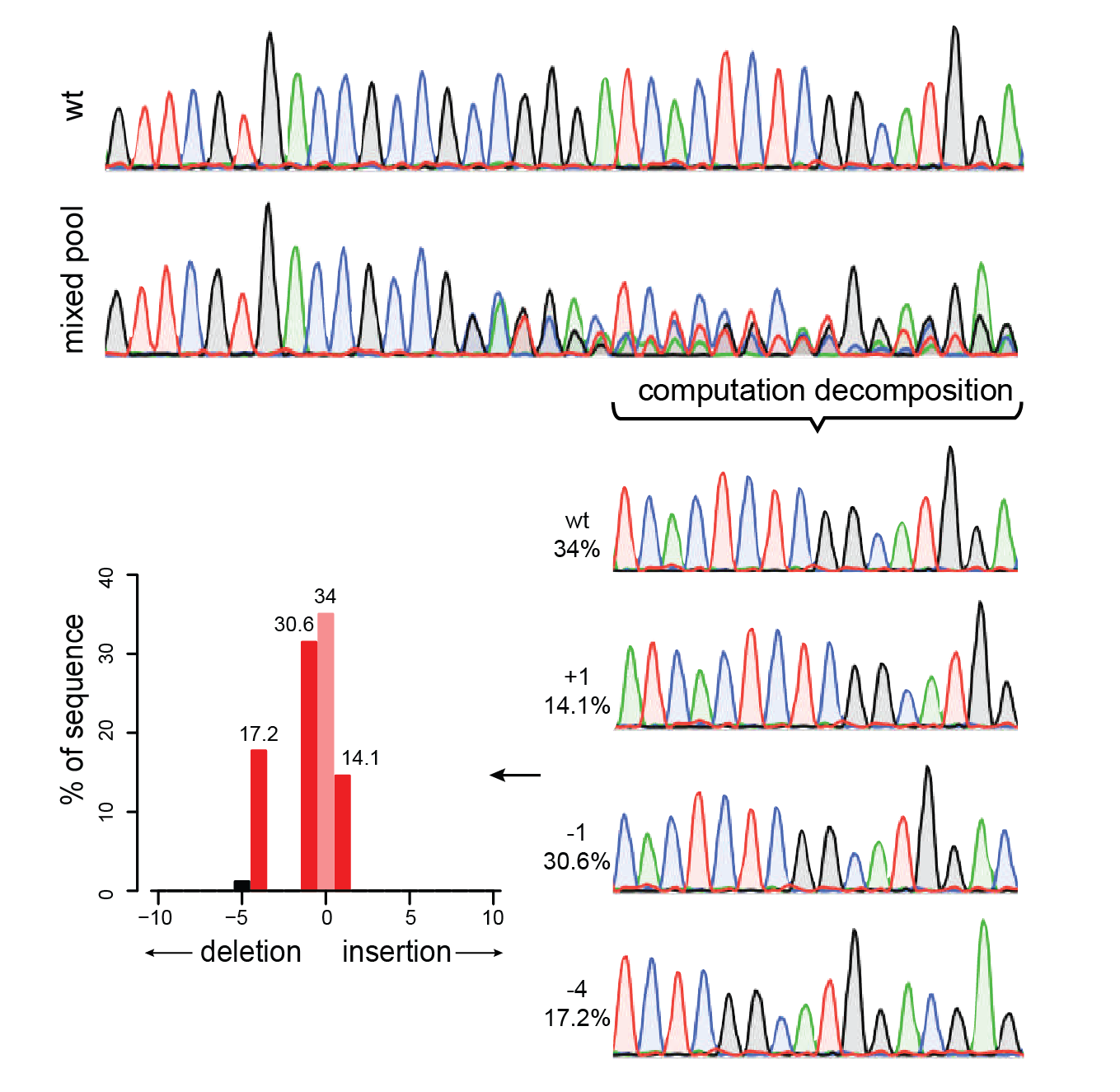
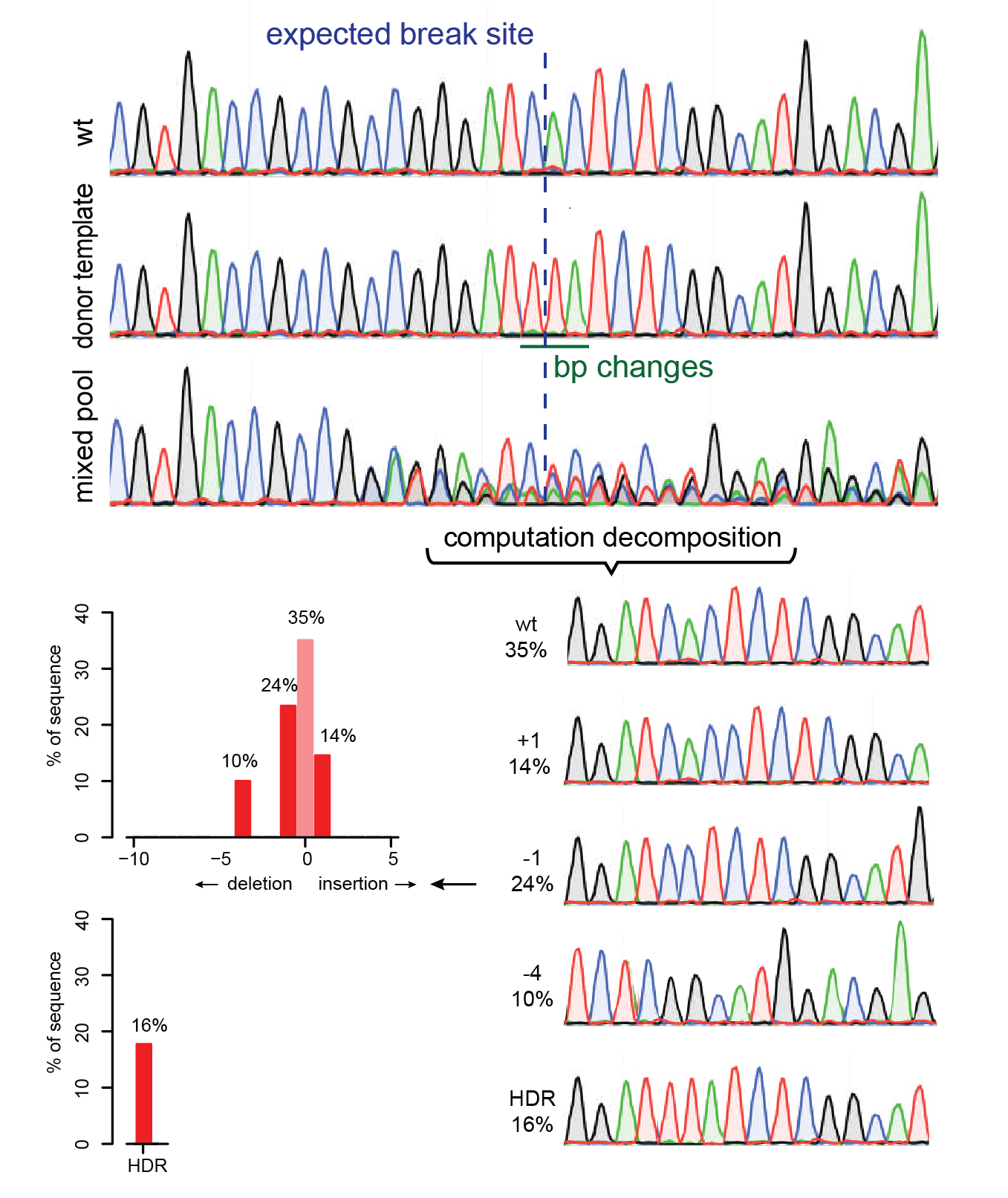
TIDER is a modified version of TIDE that estimates the frequency of targeted small nucleotide changes introduced by CRISPR in combination with
homology-directed repair using a donor template. In addition, it determines the spectrum and frequency of non-templated indels. Compared to TIDE, TIDER
requires one additional sequencing trace (i.e., three instead of two). Preparation of this third “reference” DNA can be done with a simple
two-step PCR protocol. The web tool reports the estimated frequencies of the templated mutation and of all non-templated indels. If you are only interested
in quantifying non-templated indels, we recommend the simpler TIDE method.
View TIDER publication
Start TIDER